DATA QUALITY CHECK
Earthquake Origin Time: 2019-02-10 19:00:22 (UTC) | Region: Costa_Garganica
LOCATION INGV ONT : Longitude[E] 16.364 - Latitude[N] 42.030 - Depth[km] 10.0 - Mag 3.2
ISMD Automatic Results - NOT CHECKED version
Recent Earthquakes Map Historical Earthquakes Map Citizen Macroseismic Map |
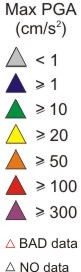
Choose view: PGA
|
|
GROUND MOTION PARAMETERS: make your stations selection and EXPORT the relative
| Station | Net | Comp. | NTC Soil | NTC Topo | PGA (cm/s^2) | PGV (cm/s) | PGD (cm) | SA03s (cm/s^2) | SA1s (cm/s^2) | SA3s (cm/s^2) | AI(cm/s) | HI (cm) | RE (km) | RY (km) | Status | Quality class | Check |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MELA | IV | HNZ | B** | T1 | 0.11176000 | 0.00139874 | 0.00022640 | 0.06358200 | 0.00624500 | 0.00230000 | 0.00018728 | 0.00362952 | 108.5 | 109.0 | Open | - | NO |
| MELA | IV | HNN | B** | T1 | 0.11616600 | 0.00286871 | 0.00021636 | 0.12448600 | 0.01362400 | 0.00174100 | 0.00013325 | 0.00576732 | 108.5 | 109.0 | Open | - | NO |
| MELA | IV | HNE | B** | T1 | 0.12264700 | 0.00270497 | 0.00017438 | 0.15522300 | 0.01481600 | 0.00195600 | 0.00026728 | 0.00621290 | 108.5 | 109.0 | Open | - | NO |
| GLNS | IT | HNZ | B* | missing | 0.10643800 | 0.00197869 | 0.00012521 | 0.06819500 | 0.00817800 | 0.00102200 | 0.00008457 | 0.00426979 | 120.2 | 120.6 | Restricted | - | NO |
| GLNS | IT | HNN | B* | missing | 0.06366000 | 0.00175248 | 0.00022069 | 0.09135200 | 0.01339600 | 0.00164400 | 0.00002269 | 0.00560903 | 120.2 | 120.6 | Restricted | - | NO |
| GLNS | IT | HNE | B* | missing | 0.08614460 | 0.00184249 | 0.00016582 | 0.11092200 | 0.01086900 | 0.00209300 | 0.00002036 | 0.00562483 | 120.2 | 120.6 | Restricted | - | NO |
| SGSC | IT | HNZ | B* | T1 | 0.01770660 | 0.00047995 | 0.00009283 | 0.04261800 | 0.00229200 | 0.00064000 | 0.00000229 | 0.00134162 | 122.4 | 122.8 | Restricted | - | NO |
| SGSC | IT | HNN | B* | T1 | 0.01546310 | 0.00048844 | 0.00006018 | 0.02573600 | 0.00322300 | 0.00061500 | 0.00000165 | 0.00146866 | 122.4 | 122.8 | Restricted | - | NO |
| SGSC | IT | HNE | B* | T1 | 0.01370570 | 0.00045590 | 0.00008362 | 0.02127700 | 0.00485200 | 0.00057000 | 0.00000166 | 0.00132605 | 122.4 | 122.8 | Restricted | - | NO |
| MOCO | IV | HNZ | B* | T1* | 0.05829350 | 0.00023598 | 0.00004496 | 0.01009000 | 0.00157900 | 0.00036700 | 0.00000593 | 0.00068839 | 124.1 | 124.5 | Open | - | NO |
| MOCO | IV | HNN | B* | T1* | 0.07938260 | 0.00033664 | 0.00003674 | 0.01877800 | 0.00272600 | 0.00054100 | 0.00000906 | 0.00092185 | 124.1 | 124.5 | Open | - | NO |
| MOCO | IV | HNE | B* | T1* | 0.13926000 | 0.00038100 | 0.00005593 | 0.01773200 | 0.00507300 | 0.00042900 | 0.00003955 | 0.00137934 | 124.1 | 124.5 | Open | - | NO |
| MCLF | IT | HNZ | B* | missing | 0.08741960 | 0.00072812 | 0.00005555 | 0.02786700 | 0.00369600 | 0.00085300 | 0.00003825 | 0.00168342 | 126.9 | 127.3 | Restricted | - | NO |
| MCLF | IT | HNN | B* | missing | 0.05715290 | 0.00094277 | 0.00017142 | 0.03449200 | 0.00378700 | 0.00152100 | 0.00002350 | 0.00235823 | 126.9 | 127.3 | Restricted | - | NO |
| MCLF | IT | HNE | B* | missing | 0.07376630 | 0.00096232 | 0.00011200 | 0.03238800 | 0.00380300 | 0.00065700 | 0.00001725 | 0.00200469 | 126.9 | 127.3 | Restricted | - | NO |
| GDFR | IT | HNZ | B* | missing | 0.03783120 | 0.00071606 | 0.00013279 | 0.03451100 | 0.00340500 | 0.00093000 | 0.00000685 | 0.00199768 | 132.0 | 132.4 | Restricted | - | NO |
| GDFR | IT | HNN | B* | missing | 0.03980910 | 0.00117035 | 0.00012117 | 0.07937300 | 0.00697200 | 0.00128600 | 0.00000637 | 0.00278488 | 132.0 | 132.4 | Restricted | - | NO |
| GDFR | IT | HNE | B* | missing | 0.04830350 | 0.00124605 | 0.00009869 | 0.05249200 | 0.00513300 | 0.00095100 | 0.00000678 | 0.00248217 | 132.0 | 132.4 | Restricted | - | NO |
| GATE | IV | HNZ | B* | T3* | 0.02223830 | 0.00043942 | 0.00007670 | 0.02029500 | 0.00234800 | 0.00066500 | 0.00000168 | 0.00119184 | 133.5 | 133.9 | Open | - | NO |
| GATE | IV | HNN | B* | T3* | 0.02119560 | 0.00081072 | 0.00016140 | 0.03868200 | 0.01004400 | 0.00138900 | 0.00000216 | 0.00282316 | 133.5 | 133.9 | Open | - | NO |
| GATE | IV | HNE | B* | T3* | 0.01666730 | 0.00068550 | 0.00007475 | 0.03542400 | 0.00413400 | 0.00090000 | 0.00000209 | 0.00198381 | 133.5 | 133.9 | Open | - | NO |
| BIOG | IV | HNZ | B* | T1* | 0.02519470 | 0.00057952 | 0.00008683 | 0.01890000 | 0.00583500 | 0.00081100 | 0.00001449 | 0.00209135 | 137.8 | 138.1 | Open | - | NO |
| BIOG | IV | HNN | B* | T1* | 0.02106730 | 0.00072612 | 0.00012731 | 0.02923700 | 0.01210600 | 0.00144000 | 0.00000706 | 0.00310155 | 137.8 | 138.1 | Open | - | NO |
| BIOG | IV | HNE | B* | T1* | 0.01825920 | 0.00061095 | 0.00012244 | 0.01873500 | 0.00884900 | 0.00180800 | 0.00001007 | 0.00287416 | 137.8 | 138.1 | Open | - | NO |
| ACER | IV | HNZ | B* | T2* | 0.05710140 | 0.00159738 | 0.00010345 | 0.07705700 | 0.00614400 | 0.00082600 | 0.00000623 | 0.00268146 | 142.6 | 142.9 | Open | - | NO |
| ACER | IV | HNN | B* | T2* | 0.04597140 | 0.00167900 | 0.00012282 | 0.08318900 | 0.00881800 | 0.00102900 | 0.00000449 | 0.00355045 | 142.6 | 142.9 | Open | - | NO |
| ACER | IV | HNE | B* | T2* | 0.04955560 | 0.00234528 | 0.00014205 | 0.20783900 | 0.01409000 | 0.00107800 | 0.00000856 | 0.00507327 | 142.6 | 142.9 | Open | - | NO |
© 2024 ISMD: INGV Strong Motion Data web portal |  CITATION | COOKIE & PRIVACY | DISCLAIMER | CREDITS | CONTACTS |